Fine-Tuning Llama 3: Enhancing Accuracy in Medical Q&A With LLMs
High-quality data is essential for tuning models to solve domain-specific problems, particularly in healthcare. Given that Large Language Models (LLMs) are trained largely on scraped data from the internet, systems that rely on them have a tendency to propagate misinformation or hallucinations due to the inherent bias in the underlying datasets.
Data collection and curation is one of the most crucial components for creating high-quality LLMs. In their paper Language Models are Few-Shot Learners paper, OpenAI describes the importance of filtering and supplementing the internet-sized Common Crawl dataset with “high-quality reference corpora.” Even after the base LLM was trained, additional fine-tuning with RLHF was incorporated to impart the conversation-style we now see in ChatGPT. Furthermore, OpenAI’s training data size increased significantly between GPT-3 and GPT-4, with some estimates of more than 40x the size of the GPT-3 dataset.
The OpenAI case illustrates the importance of high-quality data and incorporating high quality sources in robust and efficient ways. When it comes to medical information, the stakes for accuracy and reliability are even higher, as misinformation can lead to real-world health risks. The costs associated with this curation—both financial and in terms of human labor—are considerable, yet the needs are justified if we are to create safe, reliable medical AI systems.
In this article, we want to demonstrate a method of curating large datasets to reduce the cost for curating a high quality medical Q&A dataset in Label Studio and fine-tuning Llama 3 on this data. We’ll highlight how to incorporate human input throughout the process, while using LLMs to aid with text generation and automation. Let’s get started.
Fine-tuning Llama 3 for Medical Q&A
Development is inherently iterative, and the process of fine-tuning Llama 3 and curating a medical Q&A dataset follows this principle. Our method is designed to be cyclic, focusing on continuous improvement through ongoing feedback and meticulous data refinement. We have structured this process into four distinct phases, as depicted in the diagram below.
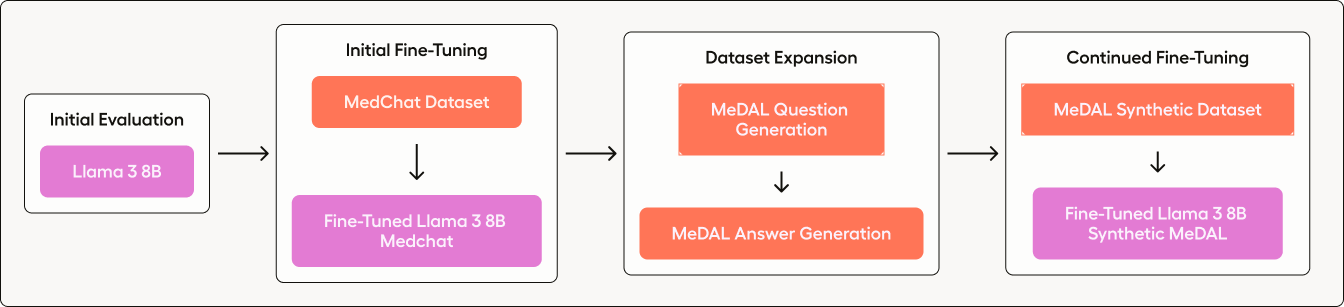
Figure 1. Diagram of the steps taken in this blog.
- Assess the Baseline Model: We start by evaluating the performance of an off-the-shelf Llama 3 8B to understand its current capabilities in handling medical queries.
- Initial Fine-Tuning: Next, we fine-tune Llama 3 using a pre-curated dataset (MedChat), to tailor the responses more closely to medical contexts.
- Dataset Expansion: We then expand our dataset by synthetically generating a large Q&A dataset from a medical diagnosis dataset (MeDAL).
- Continued Fine-tuning: Finally, we perform a second round of fine-tuning on Llama 3 using this newly curated Q&A dataset to further enhance the model’s accuracy and reliability.
Throughout these stages, we will use Label Studio's data labeling functionality to facilitate human input to modify, inspect and enhance our data. We’ll also leverage LLMs during the data curation process to not only evaluate our models results, but also make curation more effective and efficient.
We provide two Jupyter notebooks in this example: one for data curation with Label Studio and another for conducting the fine-tuning processes on a Colab T4 instance. These resources are designed to streamline the workflow and enhance the practical application of our development strategy.
- Data Curation Notebook: Utilizes Label Studio for data annotation and preparation.
- Fine-tuning Notebook: Conducts the fine-tuning processes on a Colab T4 instance, optimized for these tasks.
These resources are designed to make the development workflow more efficient and to demonstrate a practical implementation of our iterative strategy.
This structured, iterative development approach ensures that Llama 3 is not only adapted to medical Q&A but can facilitate continual improvement through systematic evaluation and refinement.
Off-the-Shelf LLMs
Open-source and freely available LLMs have impressive capabilities, but these LLMs can be very costly and time consuming to train. The most recent and impressive model in the space is Meta’s Llama 3. It performs well across many tasks, but it is a fairly general model. Let’s first investigate how it performs on a medical dataset.
The dataset that we’ll be using here is the MedChat dataset. This dataset has a variety of question-answer pairs, for testing “medical information and medical communication topics.” We should also note that like many datasets in this space, this dataset was synthetically generated, so we need to be careful with extrapolating the quality evaluation performed with such a synthetic dataset.
The first thing we’ll do is deploy Label Studio and create a project using the Label Studio SDK. This setup can be done in the UI, but we provide a code snippet to make the setup reproducible. See the Quick Start for information on deploying Label Studio.
# Styling information not displayed in this code snippet
medchat_label_config = """
<View className="root">
<Text name="chat" value="$question" layout="dialogue" />
<Header value="Answer:"/>
<Text name="summary" value="$answer" toName="summary" rows="4" editable="false" maxSubmissions="1" showSubmitButton="false"/>
<Header value="User prompt:" />
<View className="prompt">
<TextArea name="prompt" toName="chat" rows="4" editable="true" maxSubmissions="1" showSubmitButton="true" />
</View>
<Header value="Bot answer:"/>
<TextArea name="response" toName="chat" rows="4" editable="false" maxSubmissions="1" showSubmitButton="false" />
</View>
"""
# Create a new project
medchat_project = client.projects.create(
title='Project 1: MedChat',
color='#008000',
description='',
label_config=medchat_label_config
)Next, we’re going to load our dataset into Label Studio.
Our last setup step is to deploy the Label Studio Machine Learning backend with the LLM Interactive example, so we can dynamically use our LLM with our data. To avoid a lot of cloud-based infrastructure setup, we’ll use Ollama to run Llama 3 8B locally, and Docker Compose to run the LLM Backend to connect with it. Once we have these two platforms installed, we can start them with the following.
from datasets import load_dataset
medchat_dataset = load_dataset("ngram/medchat-qa")
# Create tasks for the MedChat dataset
medchat_tasks = []
for t in medchat_dataset['train']:
medchat_tasks.append({
'data': {
'question': t['question'],
'answer': t['answer']
}
})
# Import tasks into the project
for task in medchat_tasks:
client.tasks.create(
project=medchat_project.id,
data=task['data']
)Run Llama 3 locally with Ollama:
ollama run llama3To run the ML Backend, we’ll run the following:
git clone https://github.com/HumanSignal/label-studio-ml-backend.git
cd label-studio-ml-backend/label_studio_ml/examples/llm_interactive Create a docker-compose.yml file with the following configuration to connect to Ollama:
OPENAI_PROVIDER=ollama
OPENAI_MODEL=llama3
OLLAMA_ENDPOINT="http://host.docker.internal:11434/v1/" Start docker-compose.
docker-compose upWhen we navigate to Label Studio we see the project we created.
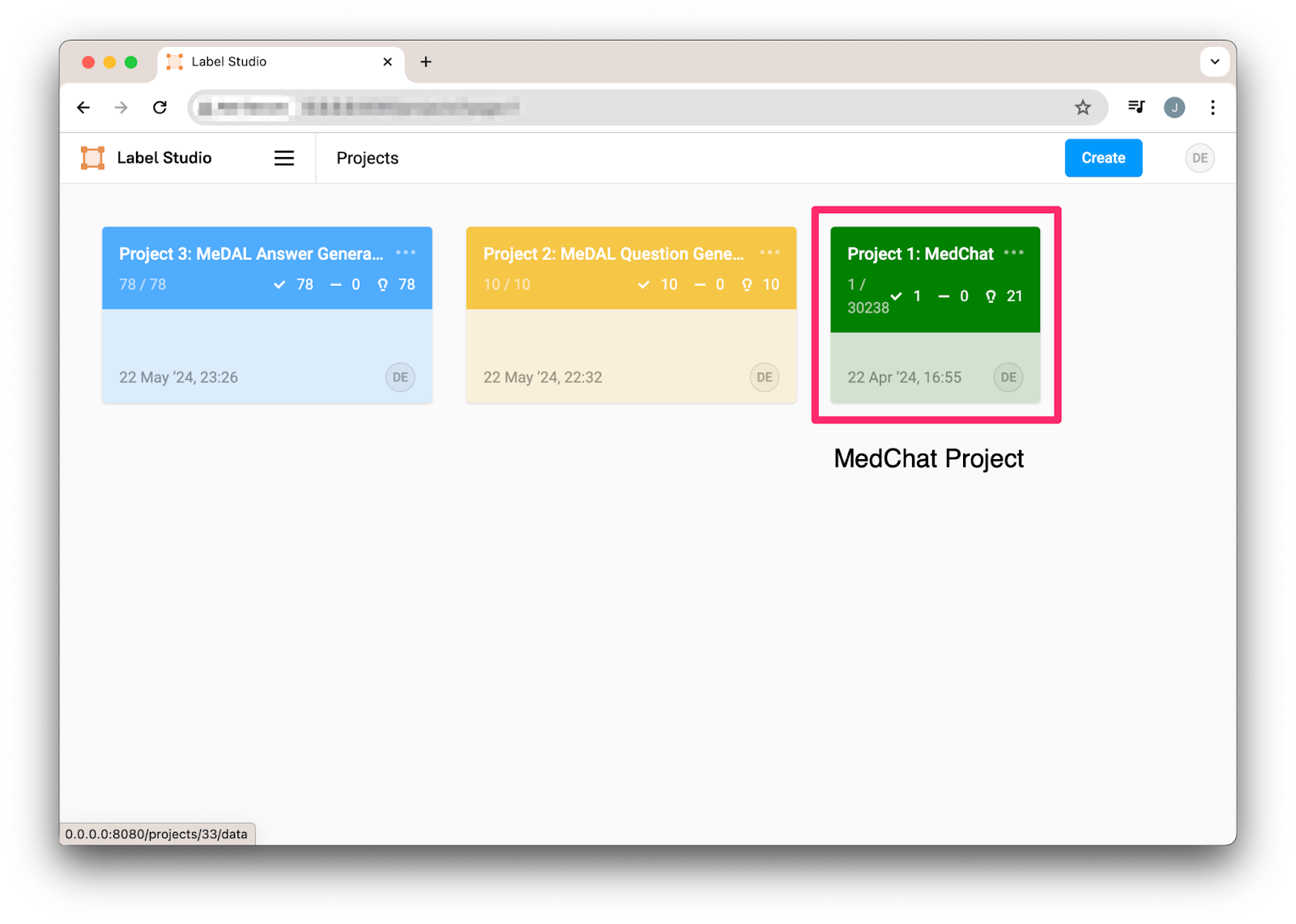
Figure 2: Med Chat project in Label Studio
To connect the ML Backend to our project, navigate to our settings, enter the address of our Docker Compose deployment, and enable “Use for interactive predictions.”
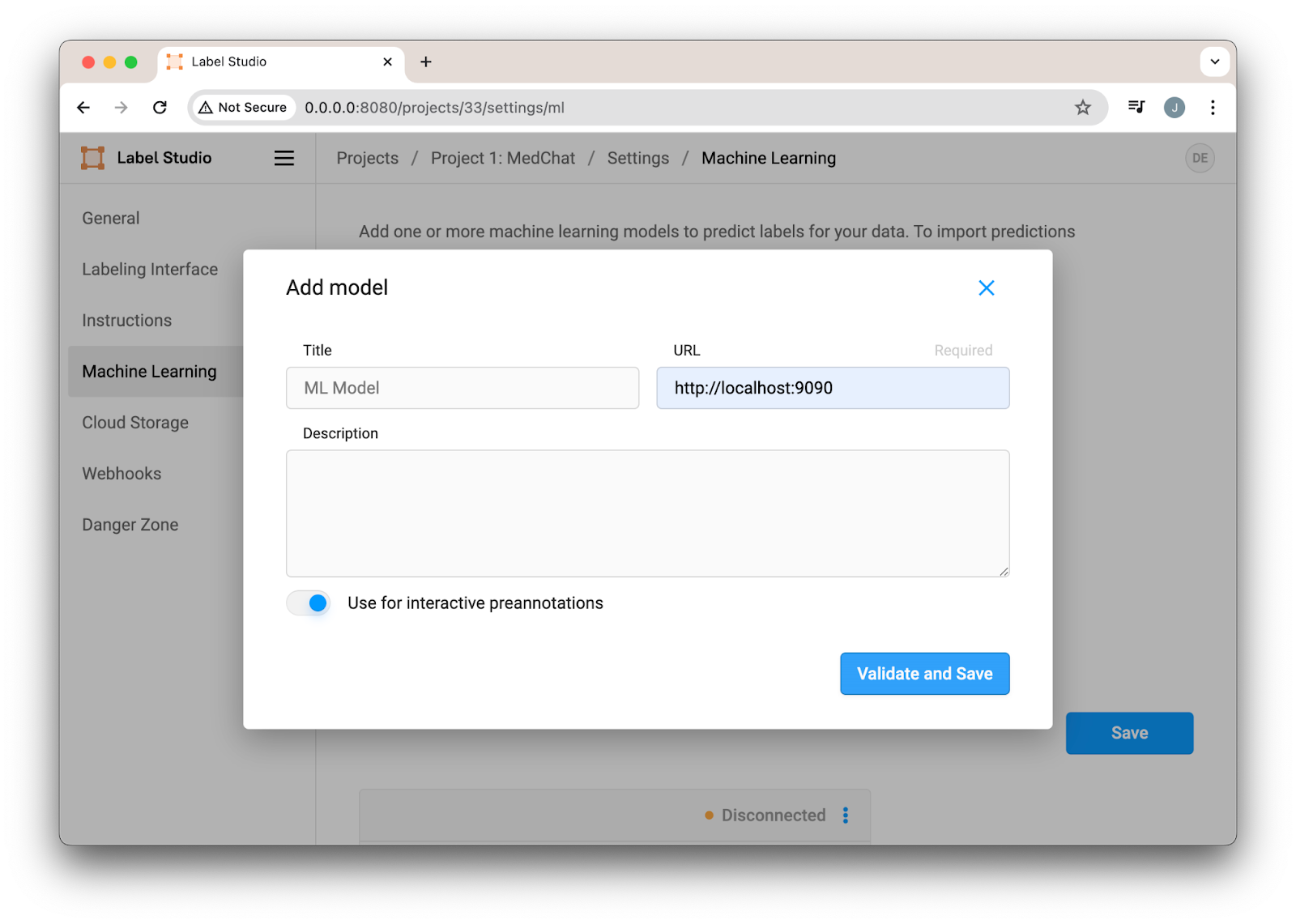
Figure 3. Configuring the Label Studio ML Backend for LLM Predictions
With the LLM Backend connected, we can use Llama 3 to propose answers to the question, and evaluate how the answer compares to the existing labeled answer. With a brief review of the existing dataset, we can already see that it has some issues that we would like to improve. For example, this Q&A pair is either missing context or is entirely wrong.

We can leverage the ML backend to make predictions using our existing model, but we should keep in mind that even if the model seems confident, we should have domain experts review it for correctness.
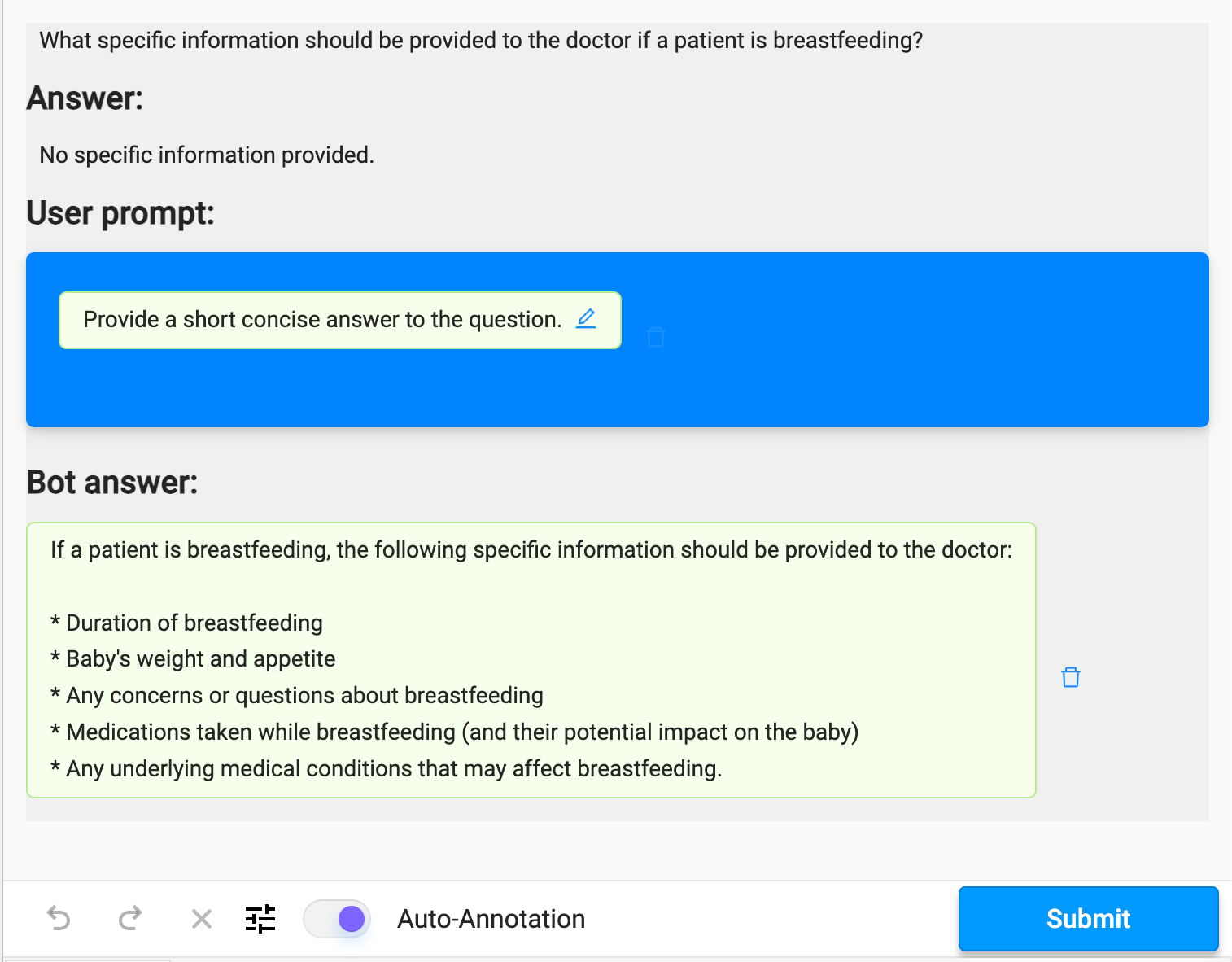
Figure 4. Evaluating the predicted answer against the LLM response.
Fine-tuning of Llama 3
We’ve seen some of the limitations of Llama 3 on the MedChat dataset. But let’s see if we can improve it by performing some fine-tuning.
Supervised fine-tuning is a process where a pre-trained model is trained (or “fine-tuned”) on a new labeled dataset. The goal of the process is to learn new insights and patterns from the new data to improve the model further on tasks that align with this dataset. In our case, we have the MedChat dataset with Q&A pairs, answering relevant medical questions. We’ll use a modified version of the Unsloth notebook we linked earlier to fine-tune our model on this dataset.
After installing our requirements, we’ll load the model that we’re going to fine-tune. In this case, we’ll use a 4-bit quantized version of Llama 3 from Unsloth, so we can train it on the free tier of Google Colab.
model, tokenizer = FastLanguageModel.from_pretrained(
model_name = "unsloth/llama-3-8b-bnb-4bit",
max_seq_length = max_seq_length,
dtype = dtype,
load_in_4bit = load_in_4bit
)Next, we’ll add our LoRA adapters. LLMs typically require large GPUs to maintain the matrices involved in the training process. To reduce the memory requirement on these GPUs, parameter-efficient fine-tuning (PEFT) techniques have emerged. One of these PEFT approaches is LoRA. LoRA takes advantage of LLM architectures in that they typically have redundancies in their weights. Therefore, LoRA trains only a small subset of the model’s parameters to save time and computational resources, which is excellent news in our resource-constrained environment. These parameters can then be merged with our LLM to produce our fine-tuned model after tuning.
model = FastLanguageModel.get_peft_model(
model,
r = 16,
target_modules = ["q_proj", "k_proj", "v_proj", "o_proj",
"gate_proj", "up_proj", "down_proj",],
lora_alpha = 16,
lora_dropout = 0,
bias = "none",
use_gradient_checkpointing = "unsloth",
random_state = 3407,
use_rslora = False,
loftq_config = None,
)With our model loaded and prepared for training, we can import our dataset. Since we haven’t made any changes to the MedChat dataset, we can import it directly from Hugging Face, provide our chat prompt template, and format our data for training. We’ll use the same process for loading our synthetic Q&A dataset later on.
chat_prompt = """
### Instruction:
{}
### Input:
{}
### Response:
{}"""
EOS_TOKEN = tokenizer.eos_token # Must add EOS_TOKEN
def formatting_prompts_func(examples):
instruction = ""
inputs = examples["question"]
outputs = examples["answer"]
texts = []
for input, output in zip(inputs, outputs):
# Must add EOS_TOKEN, otherwise your generation will go on forever!
text = chat_prompt.format(instruction, input, output) + EOS_TOKEN
texts.append(text)
return { "text" : texts, }
pass
from datasets import load_dataset
dataset = load_dataset("ngram/medchat-qa", split = "train")
dataset = dataset.map(formatting_prompts_func, batched = True,)Now we can set our training configuration and start fine-tuning (for full details, see the linked notebook).
trainer_stats = trainer.train()We can now test our model to see how it performs on new questions. Once we’re satisfied with the model, we can merge the LoRA adapters into the base model and export it in GGUF format to run in Ollama. GGUF (GPT-Generated Unified Format) is a binary format optimized for quickly loading and saving models, which makes it efficient for inference and compatible with Ollama. We can convert our model to a quantized, GGUF model with the following.
model.save_pretrained_gguf("model", tokenizer, quantization_method = "q4_k_m")Once complete, we can download our fine-tuned model and run it with Ollama. To do this, we’ll configure a model file referencing the model we downloaded.
# ./tuned-llama3-8b
FROM ./model-unsloth.Q4_K_M.gguf
TEMPLATE "{{ if .System }}<|start_header_id|>system<|end_header_id|>
{{ .System }}<|eot_id|>{{ end }}{{ if .Prompt }}<|start_header_id|>user<|end_header_id|>
{{ .Prompt }}<|eot_id|>{{ end }}<|start_header_id|>assistant<|end_header_id|>
{{ .Response }}<|eot_id|>"
PARAMETER stop <|start_header_id|>
PARAMETER stop <|end_header_id|>
PARAMETER stop <|eot_id|>
PARAMETER stop <|reserved_special_tokenWe can then import the model into Ollama.
ollama create tuned-llama3-8b -f ./tuned-llama3-8b
ollama list
ollama run tuned-llama3-8b Now to evaluate the model with the ML Backend, by modifying the model in the `docker-compose.yml` and restarting it.
- OPENAI_MODEL=tuned-llama3-8bWhen we apply this new model, we likely won’t see massive improvements in the quality of our LLM’s predictions on the dataset. They may be more succinct in areas, but overall we’ve only made minor changes through the fine-tuning process. In order to make a large improvement to the dataset, we will need a much larger, higher quality dataset. This leads us to make our own synthetic Q&A dataset with Label Studio.
Create a Synthetic Q&A Dataset
In order to create our large synthetic dataset will require a large corpus of medical knowledge. One such resource is MeDAL, a large medical text dataset curated from over 14 million articles curated from PubMed abstracts.
For synthetic Q&A generation there are two key steps: question generation and answer generation. At each step, we will want to incorporate human feedback and utilize an LLM for predictions, therefore we will create a separate project in Label Studio for each step.
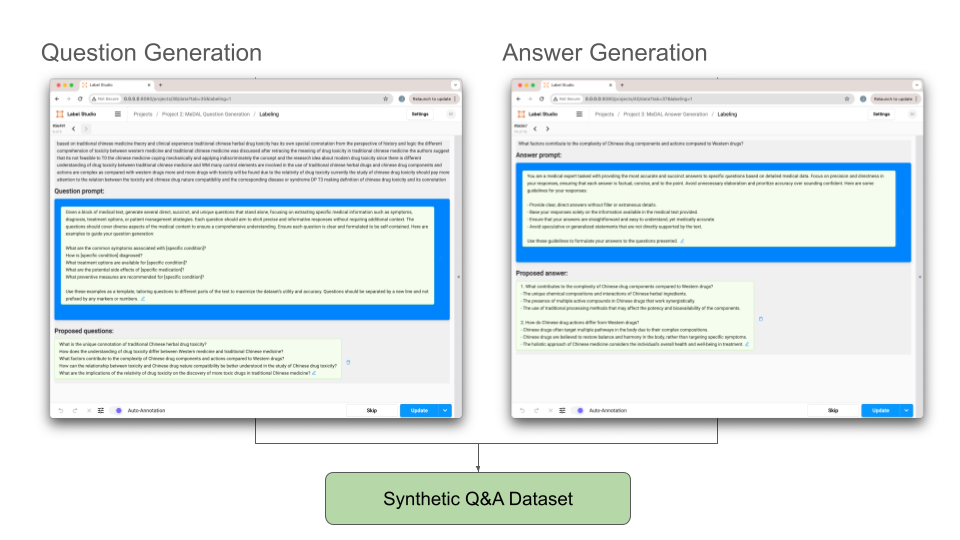
Figure 5. The outputs of the question generation and answer generation projects in Label Studio comprise our synthetic medical Q&A dataset.
First, we’ll create each of these projects by following a similar pattern to the previous sections. Notice that we still incorporate the text areas for our ML Backend to populate with predicted questions and answers.
# Styling information not displayed in this code snippet
medal_question_config = """
<View className="root">
<Text name="chat" value="$text" layout="dialogue"/>
<Header value="Question prompt:"/>
<View className="prompt">
<TextArea name="prompt" toName="chat" rows="4" editable="true" maxSubmissions="1" showSubmitButton="false"/>
</View>
<Header value="Proposed questions:"/>
<TextArea name="response" toName="chat" rows="3" editable="true" maxSubmissions="1" showSubmitButton="false"/>
</View>
"""
medal_questions_project = client.projects.create(
title='Project 2: MeDAL Question Generation',
color='#ECB800',
description='',
label_config=medal_question_config
)
medal_answer_config = '''
<View className="root">
<Text name="chat" value="$text" layout="dialogue"/>
<Header value="Answer prompt:"/>
<View className="prompt">
<TextArea name="prompt" toName="chat" rows="4" editable="true" maxSubmissions="1" showSubmitButton="false"/>
</View>
<Header value="Proposed answer:"/>
<TextArea name="response" toName="chat" rows="3" editable="true" maxSubmissions="1" showSubmitButton="false"/>
</View>
'''
medal_answers_project = client.projects.create(
title='Project 3: MeDAL Answer Generation',
color='#617ADA',
description='',
label_config=medal_answer_config
)While we could use the tuned Llama 3 model to generate our questions and answers, we can achieve a higher quality starting point for data labelers by using a better model. In this case, we can switch the ML Backend to use GPT-3.5-turbo to provide a better starting point, while also saving cost. To connect the ML Backend, we can simply modify the `docker-compose.yml` to switch from Ollama to GPT-3.5-turbo and start it again.
- OPENAI_PROVIDER=openai
- OPENAI_API_KEY=<YOUR_OPENAI_KEY>
- OPENAI_MODEL=gpt-3.5-turboNow we can configure our project to connect to the ML Backend as before and ingest our MeDAL dataset.
medal_dataset = load_dataset("medal", split='train')
num_examples = 10 # Number of examples to import
for i in range(num_examples):
task = medal_dataset[i]
client.tasks.create(
project=medal_questions_project.id,
data=task
)Now we can iterate with our LLM Prompt to provide the best questions for the context provided by the MeDAL dataset. As always, when iterating with an LLM, much of the work goes into producing a robust prompt for your task. Here is a sample prompt from which to start:
Given a block of medical text, generate several direct, succinct, and unique questions that stand alone, focusing on extracting specific medical information such as symptoms, diagnosis, treatment options, or patient management strategies. Each question should aim to elicit precise and informative responses without requiring additional context. The questions should cover diverse aspects of the medical content to ensure a comprehensive understanding. Ensure each question is clear and formulated to be self-contained. Here are examples to guide your question generation:
What are the common symptoms associated with [specific condition]?
How is [specific condition] diagnosed?
What treatment options are available for [specific condition]?
What are the potential side effects of [specific medication]?
What preventive measures are recommended for [specific condition]?
Use these examples as a template, tailoring questions to different parts of the text to maximize the dataset's utility and accuracy. Questions should be separated by a new line and not prefixed by any markers or numbers.Once we have produced a large set of questions, we can export the labeled examples from our questions project to our answers project to generate high-quality answers.
from label_studio_sdk.data_manager import Filters, Column, Type, Operator
# Create a filter for annotated tasks
filters = Filters.create(Filters.AND, [
Filters.item(
Column.completed_at,
Operator.EMPTY,
Type.Boolean,
Filters.value(False)
)
])
# Create a view in the project for the filtered tasks
view = client.views.create(
project=medal_questions_project.id,
data={
'title': 'Annotated Tasks',
'filters': filters
}
)
tab = client.views.get(id=view.id)
# Download questions from our filtered view in Label Studio
annotated_tasks = client.tasks.list(view=tab.id, fields='all')
questions_tasks = annotated_tasks.itemsAfter downloading the generated questions tasks from Label Studio, we’ll extract the desired information and format it for our answers project.
Once we have the questions extracted and formatted, we can upload them to our answers project.
# Extract questions
def extract_questions_data(questions_tasks):
data = []
for task in questions_tasks:
for result in task.annotations[0]['result']:
if result['from_name'] == 'response':
# Extract the abstract_id
abstract_id = task.data['abstract_id']
# Extract the question text and split by newlines to handle multiple questions
questions = result['value']['text'][0].split('\n')
# Store each question with its corresponding abstract_id
for question in questions:
# Check if the question is not empty and contains at least one alphanumeric character
if question.strip() and re.search('[a-zA-Z0-9]', question):
data.append({'abstract_id': abstract_id, 'text': question})
break
return data
extracted_questions_data = extract_questions_data(questions_tasks)
questions_dataset = Dataset.from_dict({'abstract_id': [item['abstract_id'] for item in extracted_questions_data],
'text': [item['text'] for item in extracted_questions_data]})# Upload the dataset to our Answers Project
for question in questions_dataset:
client.tasks.create(
project=medal_answers_project.id,
data=question
)We use the following prompt to generate our answers, although there are many improvements to be made.
You are a medical expert tasked with providing the most accurate and succinct answers to specific questions based on detailed medical data. Focus on precision and directness in your responses, ensuring that each answer is factual, concise, and to the point. Avoid unnecessary elaboration and prioritize accuracy over sounding confident. Here are some guidelines for your responses:
- Provide clear, direct answers without filler or extraneous details.
- Base your responses solely on the information available in the medical text provided.
- Ensure that your answers are straightforward and easy to understand, yet medically accurate.
- Avoid speculative or generalized statements that are not directly supported by the text.
Use these guidelines to formulate your answers to the questions presented.Finally, once we have curated our dataset, we can download our data, format it as a Hugging Face dataset, and upload it to Hugging Face to use in our fine-tuning process.
view = client.views.create(
project=medal_answers_project.id,
data={
'title': 'Annotated Tasks',
'filters': filters
}
)
tab = client.views.get(id=view.id)
answers_tasks = client.tasks.list(view=tab.id, fields='all').items# Extract questions
def extract_answers_data(answers_tasks):
data = []
for task in answers_tasks:
for result in task.annotations[0]['result']:
if result['from_name'] == 'response':
# Extract the abstract_id
abstract_id = task.data['abstract_id']
# Extract the question text and split by newlines to handle multiple questions
answer = result['value']['text'][0]
question = task.data['text']
# Store each question with its corresponding abstract_id
data.append({'abstract_id': abstract_id, 'question': question, 'answer': answer})
return data
extracted_answers_data = extract_answers_data(answers_tasks)
qa_dataset = Dataset.from_dict({'abstract_id': [item['abstract_id'] for item in extracted_answers_data],
'question': [item['question'] for item in extracted_answers_data],
'answer': [item['answer'] for item in extracted_answers_data]})qa_dataset.push_to_hub("<HF_USERNAME>/med-qa")In the end, any model we train from this dataset will directly depend on its quality, therefore we set ourselves up for allowing multiple labelers, reviewers, and domain experts to review the questions and answers for correctness. We can achieve this inside Label Studio to incrementally generate a large, high quality dataset.
Fine-tune Llama 3 on Synthetic Q&A Dataset
With our synthetic Q&A dataset pushed to Hugging Face, we can now fine-tune on the new dataset. We can follow the exact same process in the previous fine-tuning section, only changing to our new dataset with the following line.
dataset = load_dataset("<HF_USERNAME>/med-qa", split = "train")As before, we can export this model and run it with Ollama again and evaluate the model inside Label Studio.
Conclusion
This blog outlines a methodical approach to adapting Large Language Models for specialized domains. By leveraging LLMs for dataset curation and integrating human input throughout the process, we can develop high-quality medical Q&A datasets for model fine-tuning. The process of dataset curation is iterative, but it can be made more efficient with tools like Label Studio. The combination of Label Studio and tools like Ollama supports a dynamic workflow that allows for continuous refinement of models. Overall, it is essential to maintain a focus on data quality and human oversight to ensure that the advancements in AI continue to serve and improve domain-specific applications effectively.